Volume 10 - Year 2022 - Pages 01-07
DOI: 10.11159/ijtan.2022.001
Evaluation of the Culture Medium in the Synthesis of Cadmium Sulfide Nanoparticles by the Fungus Fusarium Oxysporum
J. D. A. Loa1, J. L. Pichardo-Molina2, A. Calvo-Olvera1, M. O. Hernández-Jiménez1, N. G. Rojas-Avelizapa*1
1Centro de Investigación en Ciencia Aplicada y Tecnología Avanzada, Unidad Querétaro, Instituto Politécnico Nacional,
141 Cerro Blanco, Colinas del Cimatario, Querétaro, Qro., México, 76090
2Centro de Investigaciones en Óptica A.C., 115 Loma del Bosque, Lomas del Campestre, León, Gto., México, 37150.
dloa.microbiology@gmail.com; jbautistah@ipn.mx; alexa.calvo1500@gmail.com;mhernadezj2001@alumno.ipn.mx; nrojasa@ipn.mx
Abstract - Fusarium oxysporum f. sp. lycopersici is a species reported to synthesize silver, gold, and cadmium nanoparticles when using biomass; it can produce and secrete many secondary metabolites hydrolytic enzymes involved in nanoparticle synthesis processes. However, the production of these components depends on the substrate where the fungus grows. The present work aims to evaluate the effect of the growth medium of Fusarium oxysporum in the synthesis of cadmium sulfide nanoparticles (CdS-NPs). The fungal biomass was obtained from a growth medium with nitrogenous components of higher and lower complexity (MGYP and DS respectively) and used for the synthesis of CdS-NPs using cadmium nitrate (Cd (NO3)2*4H2O) and elemental sulfur (S°). CdS-NPs exhibit two absorbance bands at 320 and 450 nm, with a fluorescence emission at 520 nm when excited with a wavelength at 365 nm. Transmission electron microscopy, energy-dispersive X-ray spectroscopy, and X-ray diffraction confirmed the presence of CdS-NPs with a mean diameter of 4.7 ± 0.7 and 4.9 ± 0.9 nm when MGYP and S media were used for biomass production, as well as atomic columns of CdS-NPs with A similar crystalline structure. However, it was also possible to appreciate fewer CdS-NPs when using the fungal biomass obtained from DS medium, which was confirmed by spectrophotometry through the absorption and emission bands at 450 and 520 nm.
Keywords: Nanoparticles, biosynthesis of nanoparticles, cadmium sulfide, Fusarium oxysporum
© Copyright 2022 Authors - This is an Open Access article published under the Creative Commons Attribution License terms. Unrestricted use, distribution, and reproduction in any medium are permitted, provided the original work is properly cited.
Date Received: 2022-12-20
Date Accepted: 2022-12-22
Date Published:2022-09-25
1. Introduction
The biosynthesis of nanoparticles (NPs) is achieved by living organisms such as bacteria, plants and fungi through organic compounds [1], enzymatic and chemical reactions by nucleophilic processes, which reduces the use of materials and harmful compounds to health and the environment [2], [3]. The biosynthesis of cadmium sulfide nanoparticles (CdS-NPs) through fungal route by extracellular processes has been investigated in filamentous fungi Pleurotus ostreatus [4], Phanerochaete chrysosporium [5], Coriolus versicolor [6], Trichoderma harzianum [7], Aspergillus versicolor [8], Aspergillus niger and Fusarium oxysporum [9], [10], since they produce the biomolecules and enzymes necessary to perform the synthesis and coating of NPs by oxide-reduction processes [11], [12].
Filamentous fungi within the genus Fusarium are good candidates for the synthesis of metal NPs due to the wide variety of primary and secondary metabolites that they produce [13], [14], as a result of the extensive ecological, biological and genetic diversity of the genus [15]. Fusarium oxysporum f. sp. lycopersici is one of the most widely studied species reported for its ability to produce different NPs when its biomass is put in contact with silver [16], [17], gold, and cadmium ions [10], [18]. Therefore, the substrate or culture medium where the fungus grows is essential for the biosynthesis of CdS-NPs [3], [14], since it is responsible not only for providing the basic requirements of the fungus growth and survival for the production of biomass, which contains the enzymes and metabolites that are involved in the processes of synthesis of NPs [19]. Thus, this study aims to evaluate the effect of growth medium of Fusarium oxysporum f. sp. lycopersici in the synthesis of CdS-NPs.
2. Materials and Methods
2.1. Production of fungal biomass and synthesis of CdS-NPs
Biomass production of Fusarium oxysporum f. sp. lycopersici was done in 1.0 L flasks with 250 mL of MGYP medium (g/L: 6; malt extract, 10; glucose, 3; yeast extract, 5; gelatin peptone) or Dextrose Sabouraud medium (DS, Cat. No. 7031) with a pre-inoculum of 1×106 spores/mL. After four days of incubation at 150 rpm and 30°C, the biomass was washed with 15 mL of sterile water and recovered by vacuum filtration [20]. The negative inorganic control (Inorganic) was performed in 250 mL flasks with 60 mL of Cd(NO3)2*4H2O 1.0 mM and S° 0.1 M, while the biological control was carried out using 6.0 g of biomass grown in MGYP (Biological MGYP) or DS (Biological DS) medium and 60 mL of sterile distilled water at 24 h, 150 rpm, and 30°C .
The synthesis of CdS-NPs was made in 250 mL flasks with 6.0 g of biomass grown in MGYP (CdS-NPs MGYP) or DS (CdS-NPs DS) medium, 60 mL of Cd(NO3)2*4H2O (1.0 mM) and S° 0.1 M at 24 h, 150 rpm, and 30°C [21]. At the end, reactions were filtered with 0.45 μm nylon membranes for physicochemical characterization.
2.2. Physicochemical characterization of CdS-NPs
Optical characterization of CdS-NPs was performed on a Genesys 10S, Thermo Scientific UV-Visible and F96Pro, Luzeren fluorometer, with spectrum readings every 1 nm from 400 to 700 nm, photomultiplexing (PMT) of 5 with an excitation at 365 nm [21]. The size, structure and composition of the CdS-NPs were analyzed by high-resolution transmission electron microscopy (HRTEM) and EDX spectroscopy on a JEOL microscope, JEM-ARM200F. The polydispersity index, hydrodynamic radius, and Z-potential of the CdS-NPs were analyzed by dynamic light scattering (DLS) ina Litesizer 500 analyzer, Anton Paar [22].
3. Results and Discussion
3.1. Optical characterization of the CdS-NPs
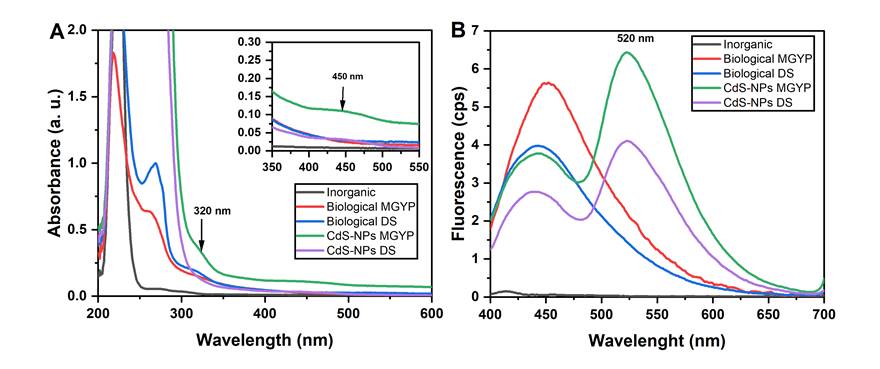
UV-Vis absorption analysis showed that inorganic precursors used in the CdS-NPs synthesis, not react by themselves, since the negative inorganic control (Inorganic) containing Cd(NO3)2*4H2O and S°, showed an absorption band at 225 nm (Fig 1A), corresponding to (NO3)22- a group of cadmium nitrate, indicating that there is no abiotic reaction between both components for CdS-NPs formation [23]. Similarly, the biological controls containing the fungal biomass in distilled water without the presence of the inorganic precursors (Biological MGYP and Biological DS) did not exhibit CdS-NPs formation, as they only showed absorption bands at 230 and 280 nm (Figure 1A), which suggest the presence of biomolecules such as carbohydrates, proteins and polar compounds secreted by the biomass [24], [25]. On the other hand, samples containing fungal biomass and inorganic precursors (CdS-NPs MGYP and CdS-NPs DS), showed an absorption band at 320 and 450 nm corresponding to the formation of CdS-NPs (Figure 1A). However, absorption was higher in the synthesis reactions when biomass was obtained from MGYP [4], [10], [21], [26].
The fluorescence analysis showed an emission band at 450 nm in Biological MGYP, Biological DS, CdS-NPs MGYP, and CdS-NPs DS (Figure 1B), which corresponds to some aromatic amino acids present in the proteins [27], as well as to the enzymatic cofactors NADH and NADPH released by the fungal biomass [28]. Nevertheless, only in CdS-NPs MGYP and CdS-NPs DS, did the fluorescence emitted at 450 nm decrease, and an emission band at 520 nm appeared (Figure 1B), which corresponds to CdS-NPs formation [29], [30]. According to previous data and based on Hietzschold results [31], we can suggest that the oxidation of NADH and NADPH cofactors occurs due to their participation in the reduction processes of metal ions to synthesize the CdS-NPs.
3.2. Morphological characterization of the CdS-NPs
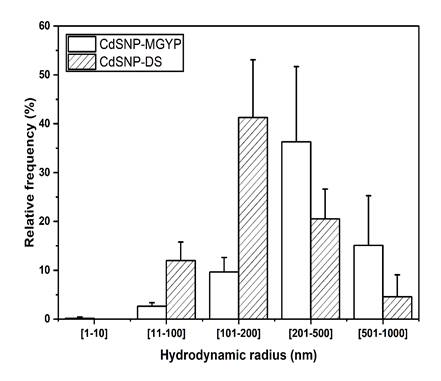
The DLS analysis showed that CdS-NPs have a wide range of hydrodynamic radius with a polydispersity index of 28%, despite the growth medium used to produce Fusarium oxysporum f. sp. lycopersici biomass (Figure 2). However, growth media (MGYP and DS) showed particles below 100 nm with a relative frequency of 2.63 ± 0.74 and 12.00 ± 3.78%, respectively. NPs below 10 nm were only observed at 0.2% of relative frequency when using the biomass produced in MGYP medium, evidencing the formation CdS-NPs. In addition, particles greater than 200 nm were also observed with a frequency higher than 20% in both culture media. It can be due to the presence of NPs clusters [22] and traces of organic material from the biomass, which is supported by the analysis of CdS-NPs at pH 5.5 by Z potential, suggesting the presence of colloidal instability of CdS-NPs MGYP and CdS-NPs DS samples which values were of 13.2 ± 1.0 mV and 9.0 ± 1.0 mV, respectively [32], [33].
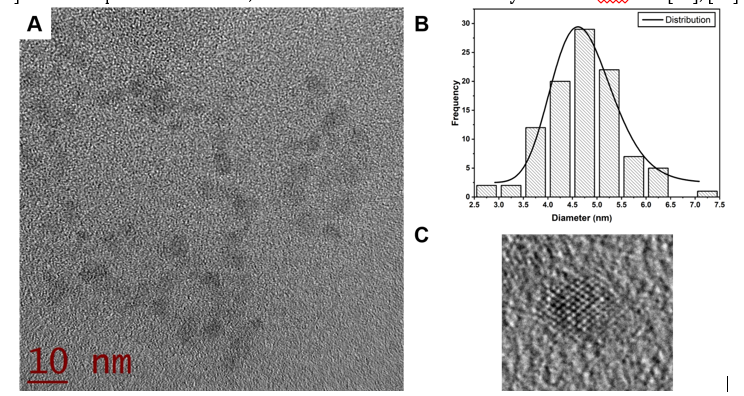
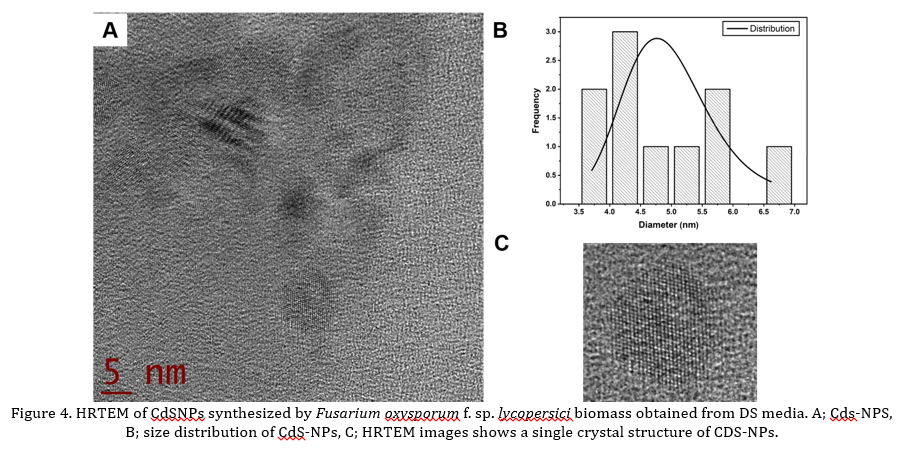
TEM micrographs show the presence of CdS-NPs (Fig 3A and Fig 4A). Also, Fig 3B and 4B show the histograms of size distribution, showing a diameter of 4.7 ± 0.7 and 4.9 ± 0.9 nm respectively, for MGYP and DS media. Besides, Fig 3C and 4C show the HRTEM images of CdS-NPs showing the crystalline planes with interplanar distances of 3.47 Å. For DS medium, fewer particles below 10 nm were observed [4], [34]. It was impossible to detect CdS-NPs by DLS, which could be attributed to the growth medium components since MGYP media contains more structurally complex components and diverse nitrogen sources than the DS medium, such as gelatin peptone, yeast extract and malt extract [35]. The composition of media, in this case MGYP, could induced a higher production and secretion of hydrolytic enzymes necessary for the formation of simple molecules from complex compounds that the fungus can capture to achieve its growth and reproduction, resulting in biomass rich in biomolecules and enzymes that help in the nucleophilic processes involved in the synthesis of CdS-NPs [36], [37].
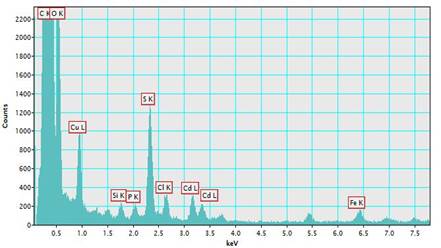
The EDX of CdS-NPs showed the presence of cadmium and sulfur, which form the CdS-NPs (Fig 5), and other elements present in the sample, like Cu, Si, and Fe, due to the TEM grid. The rest of the elements, such as C, O, and P, belong to biomolecules secreted by the fungal biomass.
4. Conclusion
The synthesis of CdS-NPs from Fusarium oxysporum f. sp. lycopersici biomass obtained in MGYP and DS medium was demonstrated by UV-Vis and fluorescence spectrophotometry, HRTEM, and DLS. It was found that the size, structure, and crystallinity of the CdS-NPs obtained are not influenced by the growth medium since no significant changes in these parameters were found. However, by using HRTEM and DLS, it was possible to appreciate that the efficiency of NPs synthesis is reduced when the fungal biomass obtained from DS medium is used since the relative frequency of CdS-NPs below 200 nm is considerably decreased. We attributed it to the organic components present in the fungal biomass, which change according to the fungus's medium.
Acknowledgements
The project was funded by the National Polytechnic Institute and Sectoral Fund for Educational Research, National Council of Science and Technology by Project A1-S-31777. Author thanks to Dr. Raúl Borja-Urby for his support with HRTEM micrographs.
References
[1] N. I. Hulkoti and T. C. Taranath, “Biosynthesis of nanoparticles using microbes-A review,” Colloids and Surfaces B: Biointerfaces, vol. 121. Elsevier, pp. 474–483, 01-Sep-2014, doi: 10.1016/j.colsurfb.2014.05.027. View Article
[2] K. S. Siddiqi and A. Husen, “Fabrication of Metal Nanoparticles from Fungi and Metal Salts: Scope and Application,” Nanoscale Research Letters, vol. 11, no. 1, pp. 1–15, 2016, doi: 10.1186/s11671-016-1311-2. View Article
[3] A. Ahmad, P. Mukherjee, D. Mandal, S. Senapati, M. I. Khan, R. Kumar, and M. Sastry, “Enzyme mediated extracellular synthesis of CdS nanoparticles by the fungus, Fusarium oxysporum,” Journal of the American Chemical Society, vol. 124, no. 41, pp. 12108–12109, 2002, doi: 10.1021/ja027296o. View Article
[4] M. Borovaya, Y. Pirko, T. Krupodorova, A. Naumenko, Y. Blume, and A. Yemets, “Biosynthesis of cadmium sulphide quantum dots by using Pleurotus ostreatus (Jacq.) P. Kumm,” Biotechnology and Biotechnological Equipment, vol. 29, no. 6, pp. 1156–1163, Nov. 2015, doi: 10.1080/13102818.2015.1064264. View Article
[5] G. Chen, B. Yi, G. Zeng, Q. Niu, M. Yan, A. Chen, J. Du, J. Huang, and Q. Zhang, “Facile green extracellular biosynthesis of CdS quantum dots by white rot fungus Phanerochaete chrysosporium,” Colloids and Surfaces B: Biointerfaces, vol. 117, pp. 199–205, 2014, doi: 10.1016/j.colsurfb.2014.02.027. View Article
[6] R. Sanghi and P. Verma, “A facile green extracellular biosynthesis of CdS nanoparticles by immobilized fungus,” Chemical Engineering Journal, vol. 155, no. 3, pp. 886–891, Dec. 2009, doi: 10.1016/j.cej.2009.08.006. View Article
[7] A. S. Bhadwal, R. M. Tripathi, R. K. Gupta, N. Kumar, R. P. Singh, and A. Shrivastav, “Biogenic synthesis and photocatalytic activity of CdS nanoparticles,” RSC Advances, vol. 4, no. 19, pp. 9484–9490, 2014, doi: 10.1039/c3ra46221h. View Article
[8] S. K. Das, I. Shome, and A. K. Guha, “Surface functionalization of Aspergillus versicolor mycelia: In situ fabrication of cadmium sulphide nanoparticles and removal of cadmium ions from aqueous solution,” RSC Advances, vol. 2, no. 7, pp. 3000–3007, Apr. 2012, doi: 10.1039/c2ra01273a. View Article
[9] M. S. Alsaggaf, A. F. Elbaz, S. El Badawy, and S. H. Moussa, “Anticancer and antibacterial activity of cadmium sulfide nanoparticles by Aspergillus niger,” Advances in Polymer Technology, vol. 2020, pp. 1–13, 2020, doi: 10.1155/2020/4909054. View Article
[10] L. R. Reyes, I. Gómez, and M. T. Garza, “Biosynthesis of cadmium sulfide nanoparticles by the Fungi Fusarium sp,” International Journal of Green Nanotechnology: Biomedicine, vol. 1, no. 1, pp. 90–95, 2009, doi: 10.1080/19430850903149936. View Article
[11] G. Sharma, S. Pandey, S. Ghatak, G. Watal, and P. K. Rai, Potential of Spectroscopic Techniques in the Characterization of “ Green Nanomaterials". Elsevier Inc., 2018. View Article
[12] N. Durán, P. D. Marcato, M. Durán, A. Yadav, A. Gade, and M. Rai, “Mechanistic aspects in the biogenic synthesis of extracellular metal nanoparticles by peptides, bacteria, fungi, and plants,” Applied Microbiology and Biotechnology, vol. 90, no. 5. Springer, pp. 1609–1624, 12-Jun-2011, doi: 10.1007/s00253-011-3249-8. View Article
[13] N. P. Keller, “Fungal secondary metabolism: regulation, function and drug discovery,” Nature Reviews Microbiology, vol. 17, no. 3, p. 167, Mar. 2019, doi: 10.1038/S41579-018-0121-1. View Article
[14] N. Durán, P. D. Marcato, O. L. Alves, G. I. H. De Souza, and E. Esposito, “Mechanistic aspects of biosynthesis of silver nanoparticles by several Fusarium oxysporum strains,” Journal of Nanobiotechnology, vol. 3, pp. 1–7, 2005, doi: 10.1186/1477-3155-3-8. View Article
[15] R. M. K. Toghueo, “Bioprospecting endophytic fungi from Fusarium genus as sources of bioactive metabolites,” Mycology, vol. 11, no. 1, p. 1, Jan. 2020, doi: 10.1080/21501203.2019.1645053. View Article
[16] M. A. Sabri, A. Umer, G. H. Awan, M. F. Hassan, and A. Hasnain, “Selection of Suitable Biological Method for the Synthesis of Silver Nanoparticles,” Nanomaterials and Nanotechnology, vol. 6, no. 1, p. 29, Jan. 2016, doi: 10.5772/62644. View Article
[17] S. A. Kumar, M. K. Abyaneh, S. W. Gosavi, S. K. Kulkarni, R. Pasricha, A. Ahmad, and M. I. Khan, “Nitrate reductase-mediated synthesis of silver nanoparticles from AgNO3,” Biotechnology Letters, vol. 29, no. 3, pp. 439–445, 2007, doi: 10.1007/s10529-006-9256-7. View Article
[18] S. Senapati, A. Ahmad, M. I. Khan, M. Sastry, and R. Kumar, “Extracellular biosynthesis of bimetallic Au-Ag alloy nanoparticles,” Small, vol. 1, no. 5, pp. 517–520, May 2005, doi: 10.1002/smll.200400053. View Article
[19] S. Srivastava, N. Pathak, and P. Srivastava, “Identification of limiting factors for the optimum growth of Fusarium oxysporum in liquid medium,” Toxicology International, vol. 18, no. 2, pp. 111–116, Apr. 2011, doi: 10.4103/0971-6580.84262. View Article
[20] I. Sandoval-Cárdenas, M. Gómez-Ramírez, N. G. Rojas-Avelizapa, and M. A. Vidales-Hurtado, “Synthesis of cadmium sulfide nanoparticles by biomass of Fusarium oxysporum f. sp. lycopersici,” Journal of Nano Research, vol. 46, pp. 179–191, 2017, doi: 10.4028/www.scientific.net/JNanoR.46.179. View Article
[21] I. Sandoval-Cárdenas, M. Gómez-Ramírez, and N. G. Rojas-Avelizapa, “Use of a sulfur waste for biosynthesis of cadmium sulfide quantum dots with Fusarium oxysporum f. sp. lycopersici,” Materials Science in Semiconductor Processing, vol. 63, pp. 33–39, Jun. 2017, doi: 10.1016/j.mssp.2017.01.017. View Article
[22] J. D. A. Loa, A. Castellanos-Angeles, L. Á. García-Tejeda, A. M. Rivas-Castillo, and N. G. Rojas-Avelizapa, “Assessment of Cadmium Sulfide Nanoparticles Synthesis by Cadmium-Tolerant Fungi,” Lecture Notes in Networks and Systems, vol. 297, pp. 145–156, Jun. 2021, doi: 10.1007/978-3-030-82064-0_12. View Article
[23] R. T. Kelly and N. G. Love, “Ultraviolet Spectrophotometric Determination of Nitrate: Detecting Nitrification Rates and Inhibition,” Water Environment Research, vol. 79, no. 7, pp. 808–812, 2007, doi: 10.2175/106143007x156682. View Article
[24] B. Raynal, P. Lenormand, B. Baron, S. Hoos, and P. England, “Quality assessment and optimization of purified protein samples: Why and how?,” Microbial Cell Factories, vol. 13, no. 1, Dec. 2010, doi: 10.1186/s12934-014-0180-6. View Article
[25] M. Taniguchi and J. S. Lindsey, “Database of Absorption and Fluorescence Spectra of >300 Common Compounds for use in PhotochemCAD,” Photochemistry and Photobiology, vol. 94, no. 2, pp. 290–327, Mar. 2018, doi: 10.1111/php.12860. View Article
[26] R. Aruna-Devi, M. Latha, S. Velumani, G. Oza, P. Reyes-Figueroa, M. Rohini, I. G. Becerril-Juarez, J. H. Lee, and J. Yi, “Synthesis and characterization of cadmium sulfide nanoparticles by chemical precipitation method,” Journal of Nanoscience and Nanotechnology, vol. 15, no. 11, pp. 8434–8439, 2015, doi: 10.1166/jnn.2015.11472. View Article
[27] J. Perucho, R. Gonzalo-Gobernado, E. Bazan, M. J. Casarejos, A. Jiménez-Escrig, M. J. Asensio, and A. S. Herranz, “Optimal excitation and emission wavelengths to analyze amino acids and optimize neurotransmitters quantification using precolumn OPA-derivatization by HPLC,” Amino Acids, vol. 47, no. 5, pp. 963–973, May 2015, doi: 10.1007/s00726-015-1925-1. View Article
[28] T. A. Giancaspero, V. Locato, and M. Barile, “A Regulatory Role of NAD Redox Status on Flavin Cofactor Homeostasis in S . cerevisiae Mitochondria,” Oxidative Medicine and Cellular Longevity, vol. 2013, p. 16, 2013, doi: 10.1155/2013/612784. View Article
[29] S. Rempel, A. D. Levin, A. Y. Sadagov, and A. A. Rempel’, “Optical properties of cadmium sulfide quantum dots in water solutions,” Physics of the Solid State, vol. 57, no. 6, pp. 1103–1107, 2015, doi: 10.1134/S106378341506027X. View Article
[30] M. Bel Haj, N. Ben Brahim, R. Mrad, M. Haouari, R. Ben Chaâbane, and M. Negrerie, “Use of MPA-capped CdS quantum dots for sensitive detection and quantification of Co2+ ions in aqueous solution,” Analytica Chimica Acta, vol. 1028, pp. 50–58, 2018, doi: 10.1016/j.aca.2018.04.041. View Article
[31] S. Hietzschold, A. Walter, C. Davis, A. A. Taylor, and L. Sepunaru, “Does Nitrate Reductase Play a Role in Silver Nanoparticle Synthesis? Evidence for NADPH as the Sole Reducing Agent,” ACS Sustainable Chemistry and Engineering, vol. 7, no. 9, pp. 8070–8076, 2019, doi: 10.1021/acssuschemeng.9b00506. View Article
[32] K. Ashok, “Experimental Methodologies for the Characterization of Nanoparticles,” in Engineered Nanoparticles, Minneapolis, MN, USA: Academic Press, 2016, pp. 125–170. View Article
[33] T. Zheng, S. Bott, and Q. Huo, “Techniques for Accurate Sizing of Gold Nanoparticles Using Dynamic Light Scattering with Particular Application to Chemical and Biological Sensing Based on Aggregate Formation,” ACS Applied Materials and Interfaces, vol. 8, no. 33, pp. 21585–21594, 2016, doi: 10.1021/acsami.6b06903. View Article
[34] N. S. Kozhevnikova, A. S. Vorokh, and A. A. Uritskaya, “Cadmium sulfide nanoparticles prepared by chemical bath deposition,” Russian Chemical Reviews, vol. 84, no. 3, pp. 225–250, 2015, doi: 10.1070/rcr4452. View Article
[35] M. C. Silva, M. C. Bertolini, and J. R. Ernandes, “Biomass production and secretion of hydrolytic enzymes are influenced by the structural complexity of the nitrogen source in Fusarium oxysporum and Aspergillus nidulans,” Journal of Basic Microbiology, vol. 41, no. 5, pp. 69–80, 2001. View Article
[36] P. Khandel and S. K. Shahi, “Mycogenic nanoparticles and their bio-prospective applications: current status and future challenges,” Journal of Nanostructure in Chemistry, vol. 8, no. 4, pp. 369–391, Dec. 2018, doi: 10.1007/s40097-018-0285-2. View Article
[37] M. Börühan Çetin, G. Ceylan, Z. Cantürk, N. Öztürk, M. K. Babayiğit, S. Yüzüak, and M. Yamaç, “Screening of Antioxidant Activity of Mycelia and Culture Liquids of Fungi from Turkey,” Microbiology, vol. 90, no. 1, pp. 133–143, 2021, doi: 10.1134/S0026261721010033. View Article

